Country Selector
Please enter a valid location
Genome Editing for Crop Improvement

Crop Insights
From Pioneer Research & Development and Agronomy Sciences - Written by Jeffry Sander, Ph.D.1, Maria Fedorova, Ph.D.2, Jesse Munkvold, Ph.D.3; and Mark Jeschke, Ph.D.4
- Key Points
- A New Era of Crop Improvement
- Genome Editing
- What is CRISPR?
- CRISPR-Cas Facilitated Crop Improvement
- Agricultural Applications of Genome Editing
- Regulatory Landscape for Genome Editing in Plants
- References
Key Points
- Genome editing is an advanced breeding tool that allows scientists to make precise, intentional changes to an organism’s DNA through addition, removal or alteration of specific genes.
- Genome editing can be used to rapidly and efficiently create improved crop varieties that are indistinguishable from those that could be obtained using traditional breeding technologies.
- The field of genome editing made a significant leap forward with the development of the CRISPR-Cas system as a gene editing tool in the early 2010s.
- Corteva Agriscience is establishing a CRISPR-Cas advanced breeding platform to develop seed products for greater environmental resiliency, productivity and sustainability.
- Corteva’s genome editing breeding platform is enabling the rapid development of crops with enhanced disease resistance and improved drought tolerance, supporting yield stability across diverse environments.
- One of the essential enablers of any innovation is the regulatory framework governing its use, and agricultural applications of genome editing are no exception.
- Science-based, risk-proportionate and globally harmonized policies for genome-edited crops are essential for translating this innovation into real-life improvements, benefiting producers and consumers globally.
A New Era of Crop Improvement
Throughout its 100-year history, Pioneer has been a leader in driving increased agricultural productivity through crop improvement. Following its founding in 1926, what was then known as the Hi-Bred Corn Company led a revolution in corn breeding that used hybridization to dramatically increase yields. With the introduction of agricultural biotechnology in the 1990s, it was demonstrated that desirable traits from non-native sources could be introduced into crops species. For example, the introduction of transgenic Bt traits from soil bacteria provided corn the ability to protect itself from damaging pests, thus improving the quantity and reliability of corn yields.
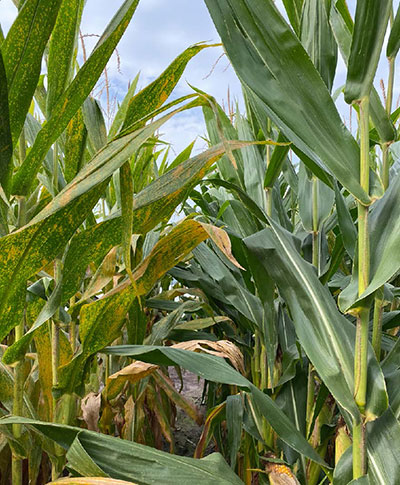

Figure 1. A gene-edited corn hybrid with multi-disease resistance (right) next to a conventional isoline hybrid (left) showing contrasting severity of southern rust (Puccinia polysora) infection. (Johnston, Iowa; September 3, 2025.)
Most recently, breakthroughs in the field of genome editing have been bringing forth a third revolution to crop improvement to be used alongside existing technologies. Genome editing is the process of introducing targeted and precise changes to DNA and other genomic features determining plant characteristics and diversity. This ground-breaking technology is expected to help develop innovative and sustainable solutions for growers similar to those realized through conventional plant breeding practices, but with even greater quality, accuracy and with more efficient development timelines.
Much of the excitement in genome editing is centered around CRISPR-Cas technology (commonly called simply CRISPR), which has been rapidly adopted due to its advantages over earlier genome editing tools in quality, efficiency and technical flexibility. CRISPR has many potential applications extending well beyond agriculture and has garnered wide mainstream media attention as research in this area has rapidly expanded. Two of the scientists who led the development of CRISPR as a gene editing tool were awarded the Nobel Prize in Chemistry in 2020 for their work – only eight years after the initial paper describing their work was published (Jinek et al., 2012), highlighting the immediate impact of their discovery.
The purpose of this Crop Insights is to provide a brief overview of what genome editing in plant breeding is, how it works and how Corteva Agriscience is using this technology to facilitate a new era of crop improvement.
Genome Editing
Genome editing is an advanced breeding tool that allows scientists to make precise, intentional changes to an organism’s DNA through addition, removal or alteration of specific genes. Gene-edited products are different from those with transgenic traits — commonly referred to as genetically modified organisms, or GMOs — because these beneficial changes mimic genetic variation found in nature and can be achieved without introducing DNA from another organism. Scientists can achieve these beneficial changes more efficiently using genome editing than other technologies that are currently available.
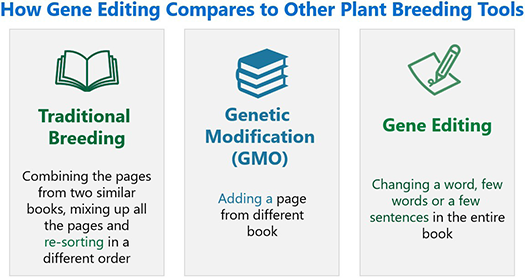

Figure 2. A useful analogy for plant breeding tools is the process of editing text in a book. Genome editing is a precise and effective way to make a beneficial change to an organism — like changing one word, a few words or a few sentences in an entire book.
While the actual implementation of genome editing is more complex, it is conceptually similar to editing a text document using a word processor. In this analogy, the genome editing tool is the cursor that can be pointed to the desired location within the text. Placement of this cursor enables one to delete, change or insert letters or even words at the selected location, thereby improving the text. In the same way a plant’s own genetic sequences can be targeted using genome editing and purposefully adapted to provide desired characteristics.
Genome editing can be used to rapidly and efficiently create improved crop varieties that are indistinguishable from those that could be obtained using traditional breeding technologies. For example, gene editing can be used to move disease-resistance alleles from lower-yielding, non-commercial genetics directly into high-yielding elite varieties. Traditional plant breeding methods can achieve the same result, but require an expensive, less precise and time-consuming backcrossing process that can take years to carry out. In contrast, genome editing preserves the integrity of the elite genetic background by introducing only the desired allele and does so within a single generation.
What is CRISPR?
The field of genome editing made a significant leap forward with the development of the CRISPR-Cas system as a gene editing tool in the early 2010s (Gasiunas et al., 2012; Jinek et al., 2012). CRISPR (clustered regularly interspaced short palindromic repeats) is a naturally occurring adaptive immune system found in many types of bacteria and archaea where it defends against viruses.
| CRISPR is a naturally occurring adaptive immune system found in many types of bacteria and archaea. |
It works by acquiring short sequences of viral DNA into the bacteria’s genome, forming a genetic memory of past invaders. When the virus attacks again, RNA transcribed from these sequences guides the Cas protein to destroy the matching viral DNA. Scientists have repurposed this system to a gene editing tool, by directing Cas enzymes to recognize and modify specific genomic sequences.
CRISPR was not the first genome editing system. The zinc-finger nuclease (ZFN) was the most widely used platform in the 2000s (Urnov et al., 2010; Gaj et al., 2016), followed by transcription activator-like effector nucleases (TALENs) in the early 2010s (Joung and Sander, 2013; Gaj et al, 2016).
However, high cost, technical complexity and limited design flexibility restricted broader adoption of these platforms. In contrast, CRISPR is more robust and significantly easier to design and use. This enables scientists to create edits more rapidly and cost-effectively and allows for the simultaneous modification of multiple genes.
Over the last decade, many CRISPR-Cas systems have been discovered and characterized including novel variants identified by scientists at Corteva Agriscience (Bigelyte et al., 2021; Urbaitis et al., 2022). Despite the diversity of these systems (e.g. sequence, structure, size, temperature) they share the common feature of a programmable RNA(s) that is capable of guiding a Cas protein to matching DNA sequences.
CRISPR-Cas Facilitated Crop Improvement
Higher organisms, including plants, continuously encounter DNA breaks from external sources such as sunlight and internal processes that release free radicals. They have developed efficient mechanisms for repairing the multitude of DNA breaks that occur in each cell every day. DNA break repairs are generally classified as non-homologous end joining or homology directed repair (Figure 3).
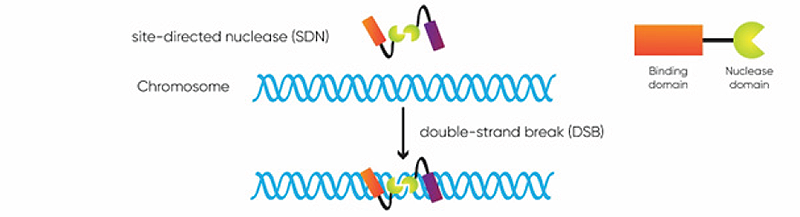
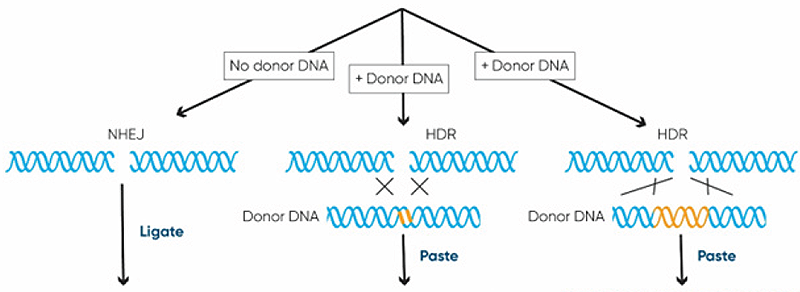

Figure 3. CRISPR-Cas facilitated DNA repair and basic CRISPR genome editing applications (from Podevin et al., 2013).
- Repair without a template occurs through the nonhomologous end joining (NHEJ) pathway and can be used to disrupt the function of a gene, effectively deleting it.
- Repair using a template through the homology-directed repair (HDR) pathway can enable precise alterations and insertions from the template DNA sequence into the genome.
- Click here or on the image above for a larger view.
Non-homologous end-joining (NHEJ) is the dominant DNA repair pathway in plants. It reconnects DNA ends introducing insertions or deletions. These changes can modulate gene expression or even turn genes off completely. Homology-directed repair (HDR) uses a second intact DNA strand, with a sequence that matches the regions flanking the break, to precisely repair it, incorporating any sequence variation between the flanking regions.
Initially, genome editing applications relied on programmable molecular scissors such as CRISPR-Cas to cut DNA at specific targets, leveraging cellular repair pathways like NHEJ and HDR to delete, modify or insert genes. More recent advances have introduced a new generation of applications capable of mimicking and mining more of nature’s diversity.
| Recent advances in genome editing have introduced a new generation of applications capable of mimicking and mining more of nature’s diversity. |
For example, Corteva scientists pioneered a method using two targeted breaks within a single chromosome to re-invert a large DNA segment that that had long been reversed relative to the standard orientation, thus restoring its ability to participate in recombination through traditional breeding (Schwartz et al., 2020). In a separate unpublished study, Corteva demonstrated the targeted relocation of a chromosomal segment from one corn chromosome to another in the same plant. These studies demonstrated that structural rearrangements such as inversions and translocations, first revealed through early sequencing of plant genomes, can be replicated using genome editing.
Next generation genome editing platforms have been expanded to go beyond double-strand breaks. New tools recruit enzymes that modify DNA without requiring double-strand breaks. For example, they can now enzymatically copy sequence from an RNA template into the genome using a single-strand nick (Anzalone et al., 2019; Ferreira da Silva et al., 2024).
| Next generation technologies are highly efficient and well-suited for multiplexing, making it possible to obtain dozens of gene edits simultaneously. |
Other approaches recruit base-editing enzymes that introduce precise sequence changes without the need to cut DNA at all (Collantes et al., 2021). These innovations are enabled by CRISPR-Cas platforms and their ability to recognize and bind specific DNA sequences with precision. These next generation technologies are highly efficient and well-suited for multiplexing, making it possible to obtain dozens of gene edits simultaneously (Figure 4).
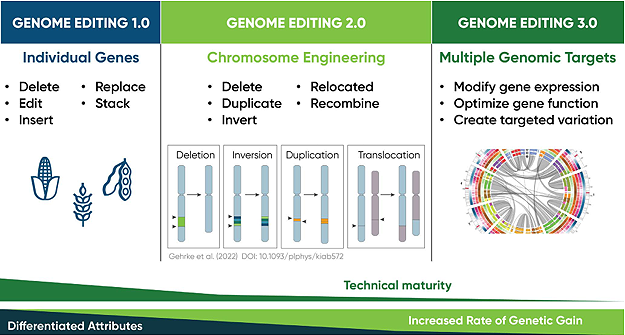
Figure 4. The next generation of genome editing technologies allow for more complex modifications like multiplexed gene activation or large-scale chromosome engineering. These newer systems are more precise and efficient, enabling scientists to edit larger sections of the genome, replace segments of DNA or activate multiple genes at once. Click here or on the image above for a larger view.
Agricultural Applications of Genome Editing
Corteva Agriscience is establishing a CRISPR-Cas advanced breeding platform to develop seed products for greater environmental resiliency, productivity and sustainability. CRISPR-Cas has numerous potential agricultural applications including improvements to yield, disease resistance and drought tolerance, as well as improvements beneficial to the end user such as output characteristics and nutritional content.
Multi-Disease Resistant Corn
Corteva is using CRISPR genome editing technology to improve the genetic resistance of corn hybrids to multiple major diseases by combining and repositioning corn disease resistance genes. Global corn genetics offer a rich source of natural disease resistance genes; however, these genes may be in varieties that are lower-yielding or not adapted to the target growing environment. Additionally, these genes are often located on different chromosomes, far apart from each other.
| Using CRISPR genome editing technology, desired native disease resistance genes are co-located together and efficiently moved into modern, high-yielding corn varieties. |
Combining the desired disease resistance genes in the same modern elite inbreds through conventional breeding is very time and resource consuming and may result in a genetic linkage drag. Using CRISPR genome editing technology, the desired native disease resistance genes are co-located together and efficiently moved into modern, high-yielding corn varieties.
Corteva’s first multi-disease resistant product will protect against gray leaf spot (Cercospora zeae-maydis), northern corn leaf blight (Exserohilum turcicum), southern corn rust (Puccinia polysora), and anthracnose stalk rot (Colletotrichum graminicola). This genome editing strategy is highly adaptable, enabling deployment across other crops, targeting additional diseases and extending to geographies worldwide.
Improved Drought Tolerance in Corn
Drought stress is the primary yield-limiting factor in corn production in most regions of the world and improved drought tolerance has long been a focus of corn breeders. Drought tolerance is a complex trait involving multiple physiological processes. Nevertheless, ethylene – a gaseous plant hormone that influences plant growth and development – is known to play an important role in modulating plant response to abiotic stress, including water deficits and high temperatures.
One of the first demonstrations of this hormone’s impact on drought tolerance in corn was via manipulation of the ethylene biosynthesis gene: ACS6. Transgenic studies showed that downregulation of this gene could reduce ethylene levels and improve grain yield under drought stress conditions (Habben et al., 2014). Another ethylene associated gene, ARGOS8, is a naturally occurring negative modulator of corn’s native ethylene response. This association led Corteva scientists to pursue increasing the expression of this gene as a means to increase drought-tolerance in corn hybrids.
| CRISPR ARGOS8 variants increased grain yield by 5 bu/A under flowering drought stress conditions and did not exhibit yield loss under well-watered conditions. |
Over 400 corn inbreds were initially evaluated for native variation of expression of the ARGOS8 gene; however, despite extensive efforts using years of traditional breeding methods, the expression levels in all these lines were less than needed to have a meaningful effect on drought tolerance. Scientists then employed CRISPR-Cas gene editing technology to increase the expression level of the ARGOS8 gene by using a promoter from another native maize gene (GOS2). Field evaluations showed that, compared to the wild type, CRISPR ARGOS8 variants increased grain yield by 5 bu/A under flowering drought stress conditions and did not exhibit yield loss under well-watered conditions (Shi et al., 2017).
Gene Editing as a Breeding Tool
Plant breeding programs heavily depend on recombination — the natural reshuffling of traits that occurs when two varieties are crossed — to identify offspring with the most favorable trait combinations. This enables the development of high-performing commercial varieties with enhanced yield and other agronomic traits. Genetic recombination is naturally limited in any new breeding cross, with on average only 1-2 DNA crossovers occurring per chromosome. This recombination can be further limited where naturally occurring DNA rearrangements have taken place within the genome of one parent but not another.
Advancements in DNA sequencing technologies have made it possible to sequence entire genomes and compare them across different varieties of the same crop. Corteva Agriscience research on corn has uncovered multiple natural instances of large-scale chromosomal rearrangements that spontaneously occur in all 10 chromosomes, such as large deletions, duplications, translocations or inversions of DNA. These types of large-scale chromosomal rearrangements are not unique to corn — they are known to occur in many plant species.
| Corteva Agriscience research on corn has uncovered multiple natural instances of large-scale chromosomal rearrangements that spontaneously occur in all 10 chromosomes, such as large deletions, duplications, translocations or inversions of DNA. |
One such spontaneous DNA rearrangement that Corteva scientists observed in certain inbred lines was a large inversion of the central part of chromosome 2. The inversion covers 75.5 Megabases of the DNA sequence, which is about one-third of the chromosome. This inversion was detected in 3 out of 66 sequenced inbred lines and happened spontaneously at some point in their breeding history. Discovery of this spontaneous inversion helped explain why breeders had not observed genetic recombination in this region after crossing these three inbreds with other inbreds.
Corteva scientists decided to test if it would be possible to ‘re-invert’ this chromosomal fragment using CRISPR-Cas genome editing technology, so that its orientation would match this region in most other inbreds. Since no genes were deleted, edited or inserted — just a fragment of chromosome inverted — there is no discrete phenotype. The effect of re-inversion had to be confirmed by analyzing if recombination of characteristics can now occur in this region. Recombination in this chromosomal region has been successfully confirmed using molecular markers (Schwartz et al., 2020), which will allow breeders to unlock useful genetic variation contained in this region.
| Corteva plant breeders can significantly increase the number of new genetic combinations to evaluate, including rare combinations that may lead to increased yield and other improved agronomic performance. |
In complementary work, Corteva scientists have also used CRISPR-Cas genome editing to increase the rate of recombination in the corn genome. Targeting genes known to modulate the frequency of recombination in plants, researchers used CRISPR-Cas to turn off these genes in corn inbreds and then measured the amount of recombination when crossed with other lines. In these crosses, recombination was increased up to five-fold with no negative effects on plant performance. By increasing genetic recombination in this way, Corteva plant breeders can significantly increase the number of new genetic combinations to evaluate, including rare combinations that may lead to increased yield and other improved agronomic performance.
Regulatory Landscape for Genome Editing in Plants
One of the essential enablers of any innovation is the regulatory framework governing its use, and agricultural applications of genome editing are no exception. Regulatory policies for biotechnology were established in various countries over 20-30 years ago, at the dawn of plant genetic engineering when it was used to insert DNA ‘foreign’ to the recipient genome (DNA from a different species, e.g. bacteria). These biotechnology policies govern cultivation, food and feed uses of transgenic crops, such as Bt corn and soybean.
| Regulatory policies for biotechnology were established in various countries over 20-30 years ago, at the dawn of plant genetic engineering. |
Most countries set their policies in alignment with The Cartagena Protocol on Biosafety — an international agreement aimed to ensure the safe handling, transport and use of living modified organisms (LMOs, commonly known as GMOs — genetically modified organisms). LMO/GMO is defined as a “living organism that possesses a novel combination of genetic material obtained through the use of modern biotechnology” (Secretariat of the Convention on Biological Diversity, 2000). Seen as inherently different from conventional (non-transgenic) crops, GMO crops require onerous safety assessment in most countries and regulatory approvals before their testing and commercial cultivation.
Crop genome editing using CRISPR is a little over ten years old and one of the most recent biotechnology innovations. It allows the introduction of targeted changes to plant’s genome without leaving any foreign (transgenic) DNA sequences. Similar genetic changes can arise in plants spontaneously or by using conventional breeding techniques (e.g., chemical or irradiation mutagenesis or genetic crosses) — and thus, can occur in conventional crops. This has presented a fundamental question: do such genome-edited plants possess a “novel combination of genetic material,” are therefore GMOs, and subject to onerous GMO regulation? Or are such genome-edited plants much more similar to, and therefore as safe as, conventional varieties?
Regulators and policy makers have been tasked with addressing very important questions. How should the existing biotechnology regulations be adapted to account for genome editing? What is the appropriate regulatory policy framework to facilitate innovation while protecting human health and the environment?
Corteva Agriscience supports the position of International Seed Federation that plant varieties developed through the latest plant breeding methods, such as CRISPR genome editing, should not be differentially regulated if they are similar to or indistinguishable from varieties that could have been produced through earlier breeding methods. It is the characteristics of the product itself, and not the tool used to create it, that determines product safety.
| It is the characteristics of the product itself, and not the tool used to create it, that determines product safety. |
Regulatory policies for genome editing in plants have made tremendous progress in the past decade. A growing number of countries consider certain outcomes of genome editing as not resulting in a “GMO” and thus, consider such genome-edited crops as being equivalent to conventional crops. Global harmonization of regulatory policies is extremely important since many crops are internationally traded agricultural commodities.
For example, offering genome-edited corn hybrids to the U.S. farmers would need to consider the regulatory status of the resulting grain in major U.S. corn export markets. Therefore, regulatory policies that are aligned between countries are essential for bringing genome editing innovations to market.
| Global harmonization of regulatory policies is extremely important since many crops are internationally traded agricultural commodities. |
The regulatory landscape for plant genome editing is continuously evolving. New examples of genome-edited crops with enhanced disease resistance, increased yields, resilience to abiotic stress or improved nutritional value constantly emerge, showcasing the value of this plant breeding innovation to help address global food security and climate change challenges. This puts pressure on the regulatory policies to keep up with technological progress, but not all governments have formulated their regulatory position yet.
The Global Farmers statement on plant breeding innovations, endorsed by 30 international farmer and agricultural industry organizations, states: “Farmers urge governments to remove regulatory impediments and uncertainty to advance plant breeding solutions for rural communities, food security and sustainable development.” Science-based, risk-proportionate and globally harmonized policies for genome-edited crops are essential for translating this scientific innovation into real-life improvements, benefiting producers and consumers globally.
References
- Anzalone, A.V., P.B. Randolph, J.R. Davis, A.A. Sousa, L.W. Koblan, J.M. Levy, P.J. Chen, C. Wilson, G.A. Newby, A. Raguram, and D.R. Liu. 2019. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature, 576(7785), 149–157.
- Bigelyte, G., J.K. Young, T. Karvelis, K. Budre, R. Zedaveinyte, V. Djukanovic, E. Van Ginkel, S. Paulraj, S. Gasior, S. Jones, L. Feigenbutz, G. St. Clair, P. Barone, J. Bohn, A. Acharya, G. Zastrow-Hayes, S. Henkel- Heinecke, A. Silanskas, R. Seidel, and V. Siksnys. 2021. Miniature type V-F CRISPR-Cas nucleases enable targeted DNA modification in cells. Nat. Comm. 12(1):6191.
- Collantes, J.C., V.M. Tan, H. Xu, M. Ruiz-Urigüen, A. Alasadi, J. Guo, H. Tao, C. Su, K.M. Tyc, T. Selmi, J.J. Lambourne, J.A. Harbottle, J. Stombaugh, J. Xing, C.M. Wiggins, and S. Jin. 2021. Development and Characterization of a Modular CRISPR and RNA Aptamer Mediated Base Editing System. CRISPR J. 4(1):58-68.
- Ferreira da Silva, J., C.J. Tou, E.M. King, M.L. Eller, D. Rufino-Ramos, L. Ma, C.R. Cromwell, J. Metovic, F.M.C. Benning, L.H. Chao, F.S. Eichler, and B.P. Kleinstiver. 2024. Click editing enables programmable genome writing using DNA polymerases and HUH endonucleases. Nat Biotechnol. 43(6):923-935.
- Gaj, T., S.J. Sirk, S.L. Shui, and J. Liu. 2016. Genome-Editing Technologies: Principles and Applications. Cold Spring Harb. Perspect. Biol. 8:a023754.
- Gasiunas, G., R. Barrangou, P. Horvath, and V. Siksnys. 2012. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl. Acad. Sci. U.S.A. 109(39);E2579-86.
- Habben, J.E., X. Bao, N.J. Bate, J.L. DeBruin, D. Dolan, D. Hasegawa, T.G. Helentjaris, H.R. Lafitte, N. Lovan, H. Mo, K. Reimann, and J.R. Schussler. 2014. Transgenic alteration of ethylene biosynthesis increases grain yield in maize under field drought-stress conditions. Plant Biotechnol. J. 12:685-693.
- Jinek, M., K. Chylinski, I. Fonfara, M. Hauer, J.A. Doudna, and E. Charpentier. 2012. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816-821.
- Joung, J.K., and J.D. Sander. 2013. TALENs: a widely applicable technology for targeted genome editing Nat. Rev. Mol. Cell Bio. 14(1):49-55.
- Podevin, N., H.V. Davies, F. Hartung, F. Nogué, and J.M. Casacuberta. 2013. Site-directed nucleases: a paradigm shift in predictable, knowledge-based plant breeding. Trends in Biotechnology, Vol. 31, No. 6.
- Schwartz, C., B. Lenderts, L. Feigenbutz, P. Barone, V. Llaca, K. Fengler, and S. Svitashev. 2020. CRISPR-Cas9-mediated 75.5-Mb inversion in maize. Nat. Plants 6(12):1427-1431.
- Secretariat of the Convention on Biological Diversity. 2000. Cartagena Protocol on Biosafety to the Convention on Biological Diversity: Text and Annexes. Montreal: CBD Secretariat. Article 3, Use of Terms.
- Shi, J., H. Gao, H. Wang, H.R. Lafitte, R.L. Archibald, M. Yang, S.M. Hakimi, H. Mo, and J.E. Habben. 2017. ARGOS8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol. J. 15:207-216.
- Urbaitis, T., G. Gasiunas, J.K. Young, Z. Hou, S. Paulraj, E. Godliauskaite, M.M. Juskeviciene, M. Stitilyte, M. Jasnauskaite, M. Mabuchi, G.B. Robb, and V. Siksnys. 2022. A new family of CRISPR-type V nucleases with C-rich PAM recognition. EMBO Rep. 23(12):e55481.
- Urnov F.D., E.J. Rebar, M.C. Holmes, H.S. Zhang, and P.D. Gregory. 2010. Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 11(9):636–646.
1Jeffry Sander, Ph.D., Program Manager, Genome Editing Technologies
2Maria Fedorova, Ph.D., Global Genome Editing Technical Lead
3Jesse Munkvold, Ph.D., Program Leader, Gene Edited Breeding
4Mark Jeschke, Ph.D., Agronomy Manager
The foregoing is provided for informational use only. Contact your Pioneer sales professional for information and suggestions specific to your operation. Product performance is variable and subject to any number of environmental, disease, and pest pressures. Individual results may vary. Pioneer® brand products are provided subject to the terms and conditions of purchase which are part of the labeling and purchase documents.
